Some molecular cytogenetic markers and classical chromosomal features of Spilopelia chinensis (Scopoli, 1786) and Tachybaptus ruficollis (Pallas, 1764) in Thailand
DOI:
https://doi.org/10.36253/caryologia-952Keywords:
Streptopelia chinensis, Tachybaptus ruficollis, Bird chromosome, Bird karyotypeAbstract
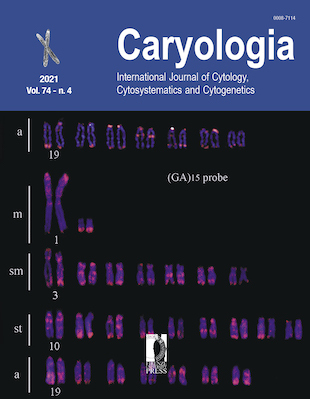
This study analyzed the karyological features of two bird species – Spilopelia chinensis and Tachybaptus ruficollis – from Northeastern Thailand. Mitotic chromosomes were indirectly prepared by fibroblast cell culture. The chromosomes were stained by conventional Giemsa staining and microsatellite repeat of fluorescence in situ hybridization techniques. Giemsa staining showed that the diploid chromosome number of S. chinensis was 2n=70 and T. ruficollis was 60. The types of chromosomes observed in S. chinensis were 4 large metacentric, 2 medium acrocentric, 2 small metacentric, 2 small submetacentric, 2 sex chromosomes and 58 microchromosomes; the karyotype of T. ruficollis comprised 2 large metacentric, 2 large submetacentric, 2 large acrocentric, 8 small metacentric, 4 small submetacentric, ZW sex chromosomes and 40 microchromosomes. The molecular cytogenetical features that were exhibited only on the male T. ruficollis chromosome included two microsatellites and telomeric sequences: two signals of d(CA)15 on two microchromosomes, one signal of d(GC)15 on one of the first pair, and signals of AGGGTTn sequences on each telomeric region of all macro- and microchromosomes. The karyotype formula was deduced as: 2n (70) = Lm4 + Ma2 + Sm2 + Ssm2 + 2 sex chromosomes (Sm1/Ssm1) + 58 microchromosomes for S. chinensis and 2n (60) = Lm2 +Lsm2 + La2 + Sm8 + Ssm4 + Z (Msm1) W (Ssm1) + 40 microchromosomes for T. ruficollis.
Downloads
References
BirdLife International, Species factsheet: Tachybaptus ruficollis. 2020. Cambridge: BirdLife International; [accessed 2020 May 25]. http://www.birdlife.org/.
Burt DW. 2002. Origin and evolution of avian microchromosomes. Cytogenet Genome Res. 96(1-4):97–112.
Chaiyasut K. 1989. Cytogenetics and cytotaxonomy of the genus Zephyranthes. Bangkok: Department of Botany, Faculty of Science, Chulalongkorn University. Thai.
Cox J, James FC. 1984. Karyotypic uniformity in the red-winged blackbird. Condor. 86:416–422.
de Oliveira EH, Tagliarini MM, dos Santos MS, O'Brien PC, Ferguson-Smith MA. 2013. Chromosome painting in three species of Buteoninae: a cytogenetic signature reinforces the monophyly of South American species. PLoS One. 8(7):e70071.
Ebied AM, Hassan HA, Abu Almaaty AH, Yaseen AE. 2005. Karyotypic characterization of ten species of birds. Cytologia. 70(2):181–194.
Ellegren H. 2000. Evolution of the avian sex chromosomes and their role in sex determination. Trends Ecol Evol. 15(5):188–192.
Getlekha N, Molina WF, Cioffi MB, Yano CF, Maneechot N, Bertollo LNC, Supiwong W, Tanomtong A. 2016. Repetitive DNAs highlight the role of chromosomal fusions in the karyotype evolution of Dascyllus species (Pomacentridae, Perciformes). Genetica. 144(2):203–211.
Gibbs D, Barnes E, Cox J. 2001. Pigeons and doves: a guide to the pigeons and doves of the world. Connecticut: Yale University Press.
Ichikawa Y, Nishimura Y, Kurumizaka H, Shimizu M. 2015. Nucleosome organization and chromatin dynamics in telomeres. Biomol Concepts. 6(1):67–75.
Jarvis ED, Mirarab S, Aberer AJ, Li B, Houde P, Li C, Ho SYW, Faircloth BC, Nabholz B, Howard JT, et al. 2014. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science. 346(6215):1320–1331.
Kaewmad P, Tanomtong A, Gomontean B, Wonkaonoi W, Khunsook S, Sanoamuang L. 2013. First karyological analysis of black crowned crane (Balearica pavonina) and scaly breasted munia (Lonchura punctulata) by conventional staining technique. Cytologia. 78(3):205–211.
Kretschmer R, Gunski RJ, Del Valle Garnero A, de Oliveira Furo I, O'Brien PCM, Ferguson-Smith MA, de Oliveira EHC. 2014. Molecular cytogenetic characterization of multiple intrachromosomal rearrangements in two representatives of the genus Turdus (Turdidae, Passeriformes). PLoS One. 9(7):e103338.
Ksepka DT, Boyd CA. 2012. Quantifying historical trends in the completeness of the fossil record and the contributing factors: an example using Aves. Paleobiology. 38(1):112–125.
Kumar R. 2018. Microsatellite marker. In: Vonk J, Shackelford T. (eds) Encyclopedia of Animal Cognition and Behavior. Cham: Springer.
Makino S, Udagama T, Yamashina Y. 1956. Karyotype studies in birds. 2: a comparative study of chromosomes in the Columbidae. Caryologia. 8(2):275–293.
Pacheco MA, Battistuzzi FU, Lentino M, Aguilar RF, Kumar S, Escalante AA. 2011. Evolution of modern birds revealed by mitogenomics: timing the radiation and origin of major orders. Mol Biol Evol. 28(6):1927–1942.
Patawang I, Tanomtong A, Getlekha N, Phimphan S, Pinthong K, Neeratanaphan L. 2017. Standardized karyotype and idiogram of Bengal monitor lizard, Varanus bengalensis (Squamata, Varanidae). Cytologia. 82(1):75–82.
Phimphan S, Tanomtong A, Chuaynkern Y, Pratumtong D. 2015. Karyological analysis of red jungle fowl (Gallus gallus gallus Linnaeus, 1758) using egg fibroblastic cell culture. KKU Sci J. 43(1):39–48. Thai.
Pratumthong D, Thunhikorn S, Duengkae P. 2011. A checklist of the birds in Thailand. Journal of Wildlife in Thailand. 18(1):152–319. Thai.
Seabury CM, Dowd SE, Seabury PM, Raudsepp T, Brightsmith DJ, Liboriussen P, Halley Y, Fisher CA, Owens E, Viswanathan G, et al. 2013. A multi-platform draft de novo genome assembly and comparative analysis for the Scarlet Macaw (Ara macao). PLoS One. 8(5):e62415.
Shibusawa M, Nishibori M, Nishida-Umehara C, Tsudzuki M, Masabanda J, Griffin DK, Matsuda Y. 2004. Karyotypic evolution in the Galliformes: an examination of the process of karyotypic evolution by comparison of the molecular cytogenetic findings with the molecular phylogeny. Cytogenet Genome Res. 106(1):111–9.
Skinner BM, Robertson L BW, Tempest HG, Langley EJ, Ioannou D, Fowler KE, Crooijmans R PMA, Hall AD, Griffin DK, Völker M. 2009. Comparative genomics in chicken and Pekin duck using FISH mapping and microarray analysis. BMC Genomics. 10:357.
Srivastava MDL, Misra M. 1971. Somatic chromosomes of Streptopdia decaocto (Golumbiformes). J Hered. 62(6):373–374.
Tange M, Nakahara K. 1938-1939. On the chromosomes of the Ring Dove, Streptopelia risoria. Okajimas Folia Anat Jpn. 17(5):477–478.
Vieira MLC, Santini L, Diniz AL, de Freitas Munhoz C. 2016. Microsatellite markers: what they mean and why they are so useful. Genet Mol Biol. 39(3):312–328.
You-Sheng R, Zhang-Feng W, Xue-Wen C, Huai-Yu Z. 2008. Study on karyotype of Streptopelia chinensis and Columba livia domestica. J Nanchang Normal Univ. 3:31–33.
Yuri T, Kimball RT, Harshman J, Bowie RCK, Braun MJ, Chojnowski JL, Han KL, Hackett SJ, Huddleston CJ, Moore WS, et al. 2013. Parsimony and model-based analyses of indels in avian nuclear genes reveal congruent and incongruent phylogenetic signals. Biol(Basel). 2:419–444.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2021 Isara Patawang, Sarawut Kaewsri, Sitthisak Jantarat, Praween Supanuam, Sarun Jumrusthanasan, Alongklod Tanomtong

This work is licensed under a Creative Commons Attribution 4.0 International License.
- Copyright on any open access article in a journal published byCaryologia is retained by the author(s).
- Authors grant Caryologia a license to publish the article and identify itself as the original publisher.
- Authors also grant any third party the right to use the article freely as long as its integrity is maintained and its original authors, citation details and publisher are identified.
- The Creative Commons Attribution License 4.0 formalizes these and other terms and conditions of publishing articles.
- In accordance with our Open Data policy, the Creative Commons CC0 1.0 Public Domain Dedication waiver applies to all published data in Caryologia open access articles.